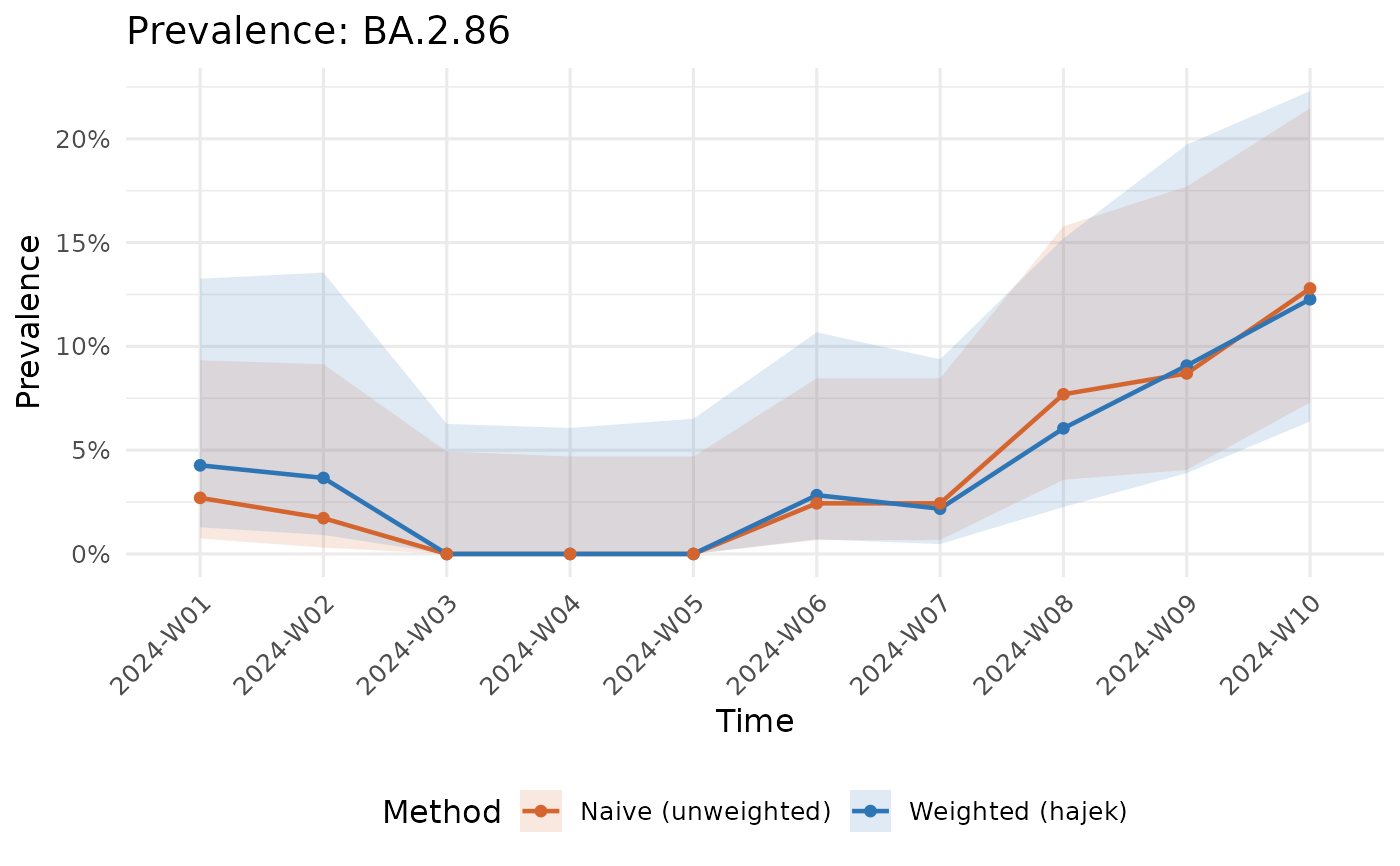
Side-by-side plot showing the impact of design correction.
Examples
sim <- surv_simulate(n_regions = 3, n_weeks = 10, seed = 1)
d <- surv_design(sim$sequences, ~ region,
sim$population[c("region", "seq_rate")], sim$population)
w <- surv_lineage_prevalence(d, "BA.2.86")
n <- surv_naive_prevalence(d, "BA.2.86")
surv_compare_estimates(w, n)