Lineage Frequency Dynamics and Growth-Advantage Estimation from Genomic Surveillance Counts
lineagefreq provides a unified pipeline for modelling pathogen lineage frequencies from genomic surveillance counts. The package implements multinomial logistic regression and alternative estimation engines, probabilistic forecasting with configurable prediction horizons, and integrated calibration diagnostics based on probability integral transform (PIT) histograms and distribution-free conformal prediction intervals. Additional modules support immune-aware fitness decomposition and information-theoretic surveillance optimisation.
Installation
# Stable release from CRAN
install.packages("lineagefreq")
# Development version from GitHub (recommended for full feature set)
# install.packages("pak")
pak::pak("CuiweiG/lineagefreq")Note: The current CRAN release (v0.2.0) provides core modelling, forecasting, and backtesting functionality. The development version on GitHub (v0.5.1) adds calibration diagnostics, conformal prediction, immune-aware fitness decomposition, and surveillance optimisation.
Quick example
library(lineagefreq)
library(ggplot2)
data(cdc_sarscov2_jn1)
x <- lfq_data(cdc_sarscov2_jn1,
lineage = lineage, date = date, count = count)
fit <- fit_model(x, engine = "mlr")
growth_advantage(fit, type = "relative_Rt", generation_time = 5)
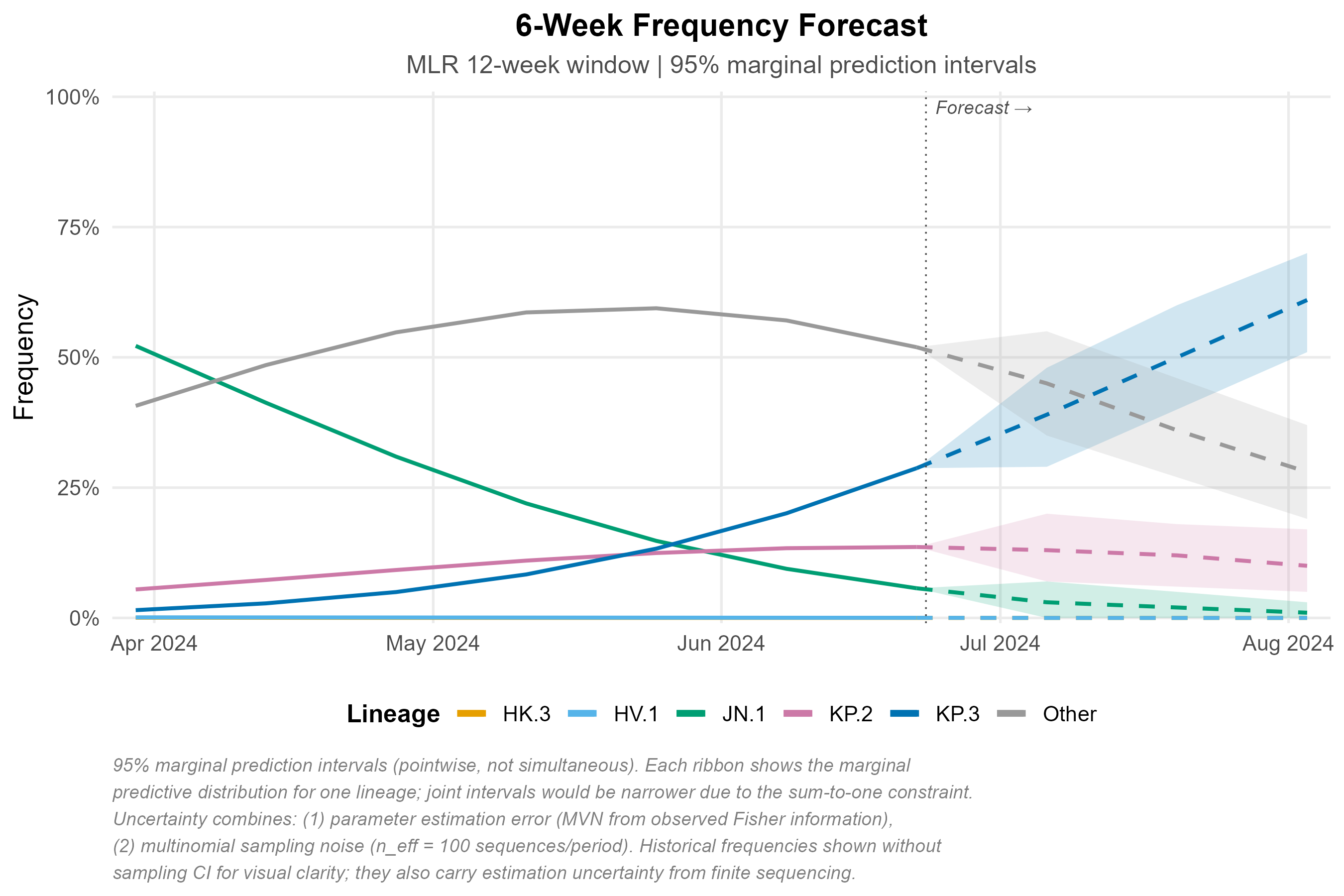
fc <- forecast(fit, horizon = 28)
autoplot(fc)Real-data case studies
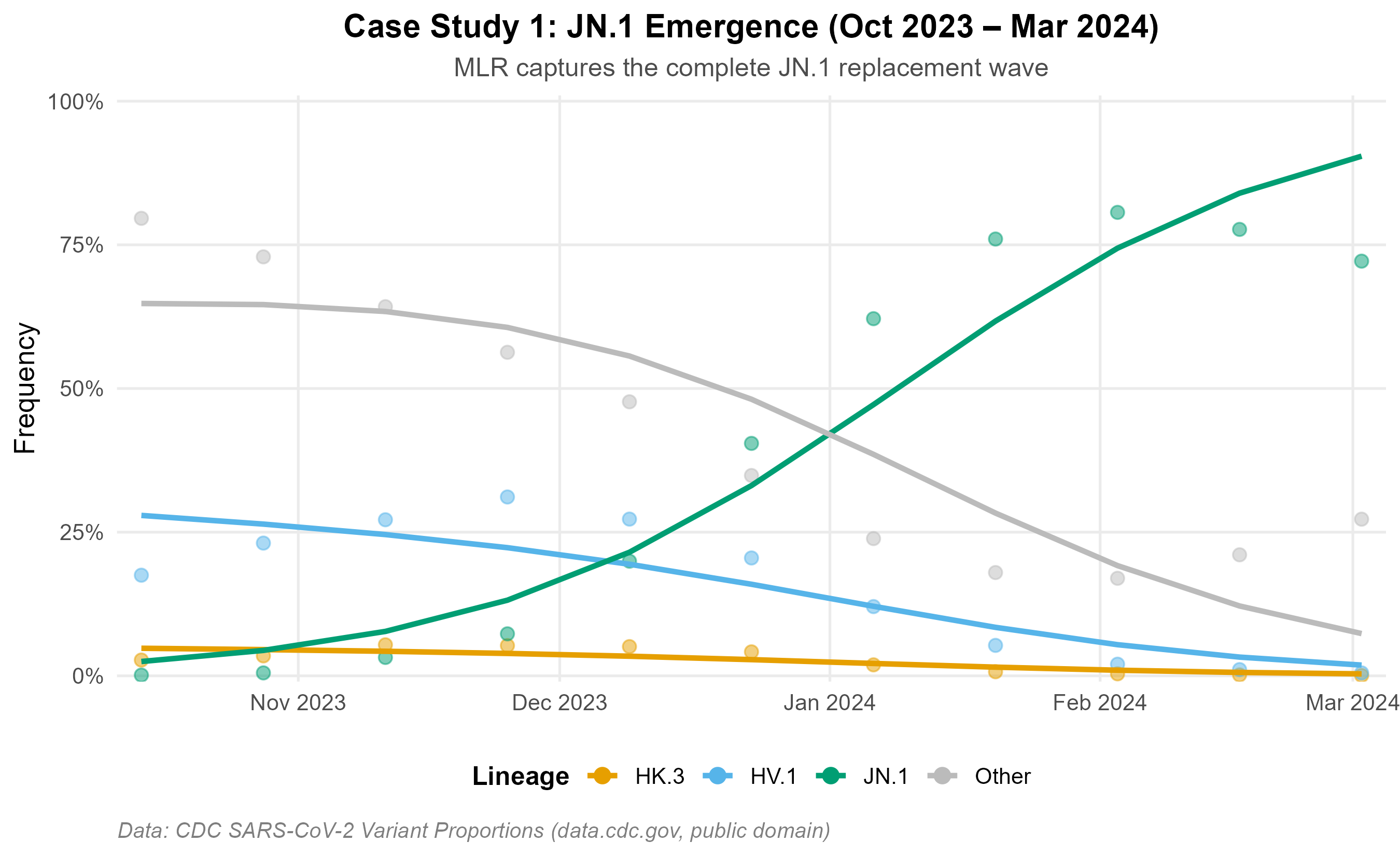
Figures below use real U.S. CDC surveillance data (data.cdc.gov/jr58-6ysp, public domain). Two independent epidemic waves illustrate model behaviour across distinct replacement settings.
Data accessed 2026-03-28. Lineages below 5% peak frequency collapsed to “Other.” Reproducible scripts: data-raw/prepare_cdc_data.R and data-raw/prepare_ba2_data.R.
Variant replacement dynamics
JN.1 emergence (Oct 2023 – Mar 2024): MLR recovers the observed replacement trajectory from <1% to >80%.

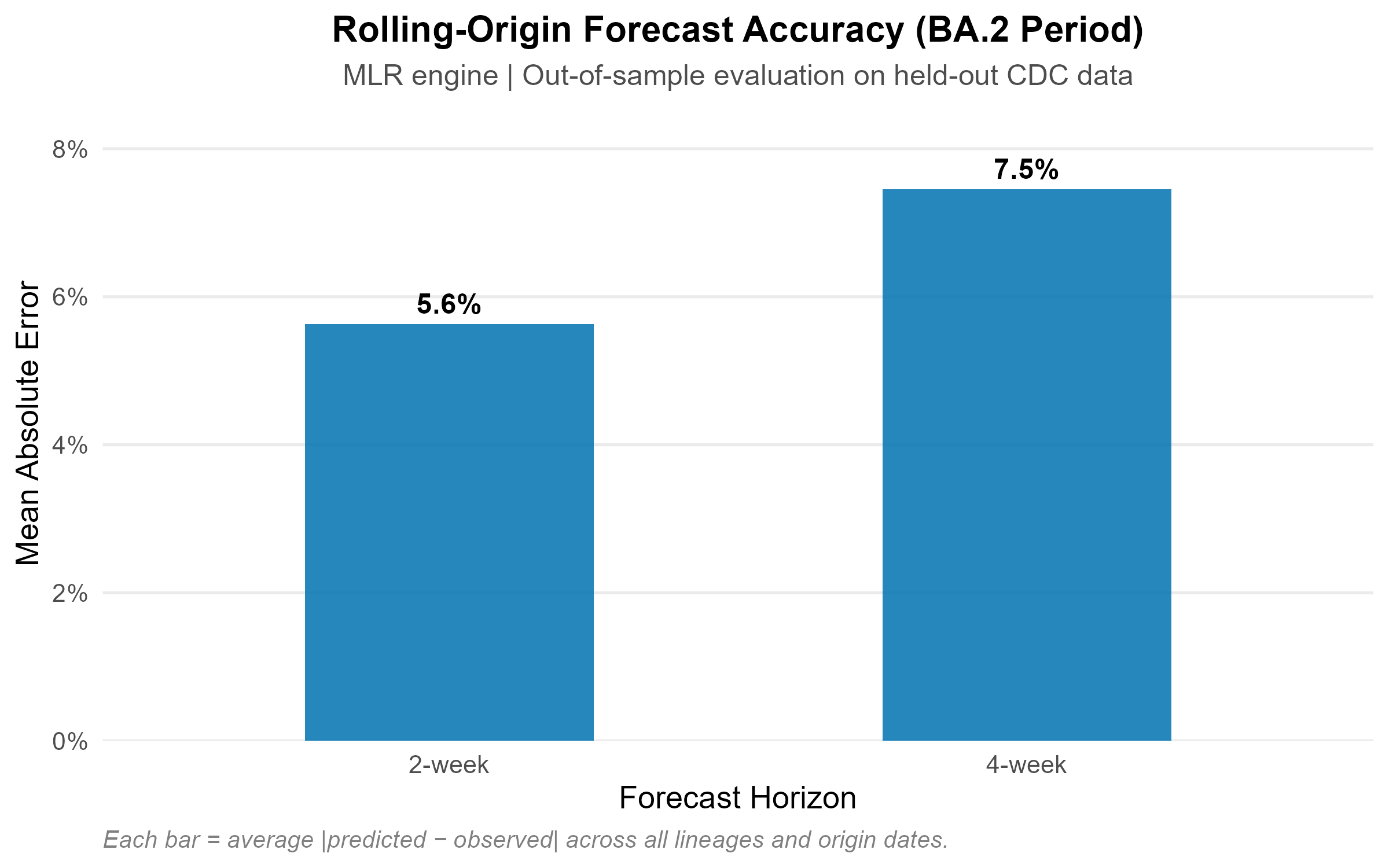
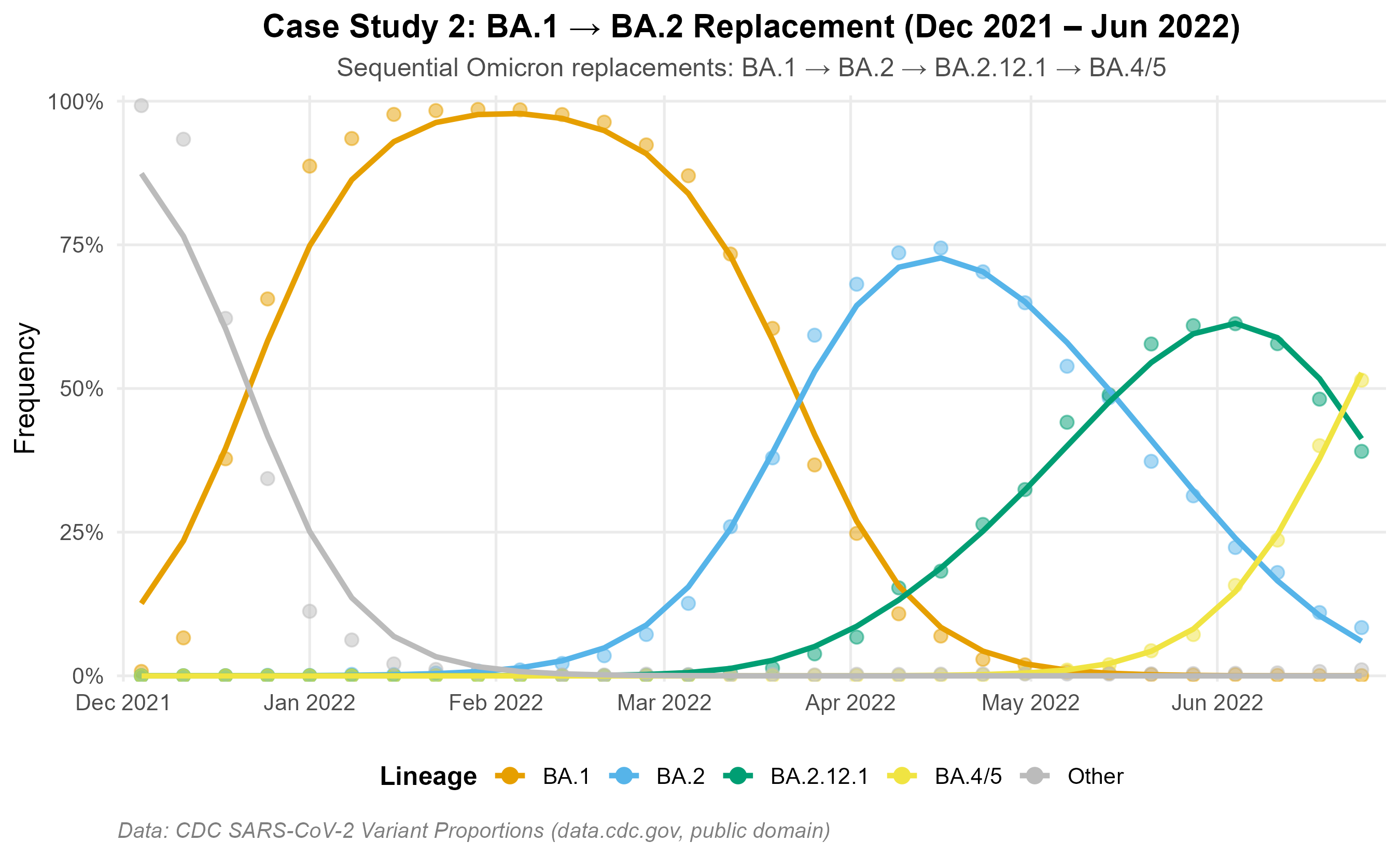
BA.1 to BA.2 period (Dec 2021 – Jun 2022): A well-characterised Omicron replacement wave with four sequential subvariant sweeps.
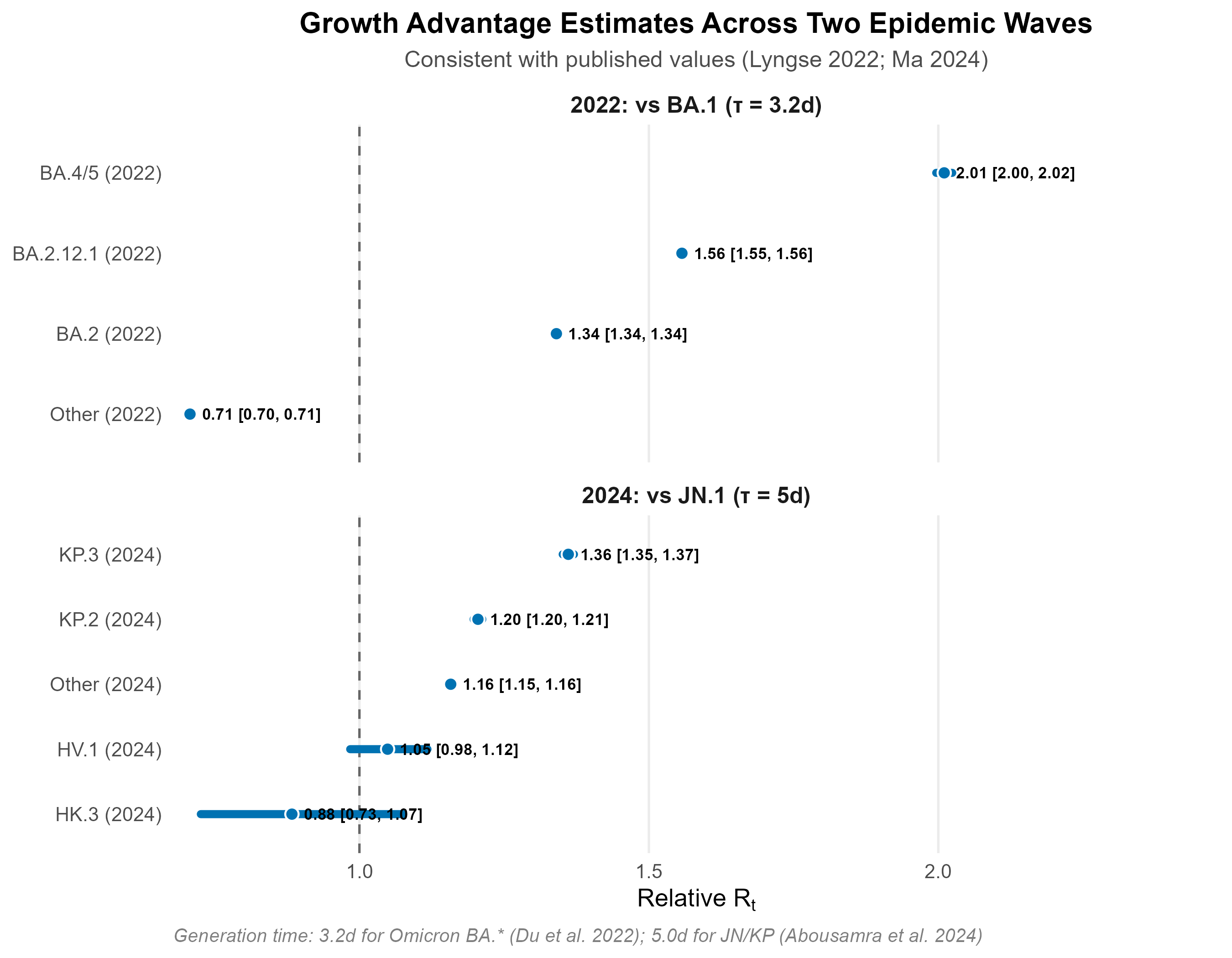
Growth advantage estimation
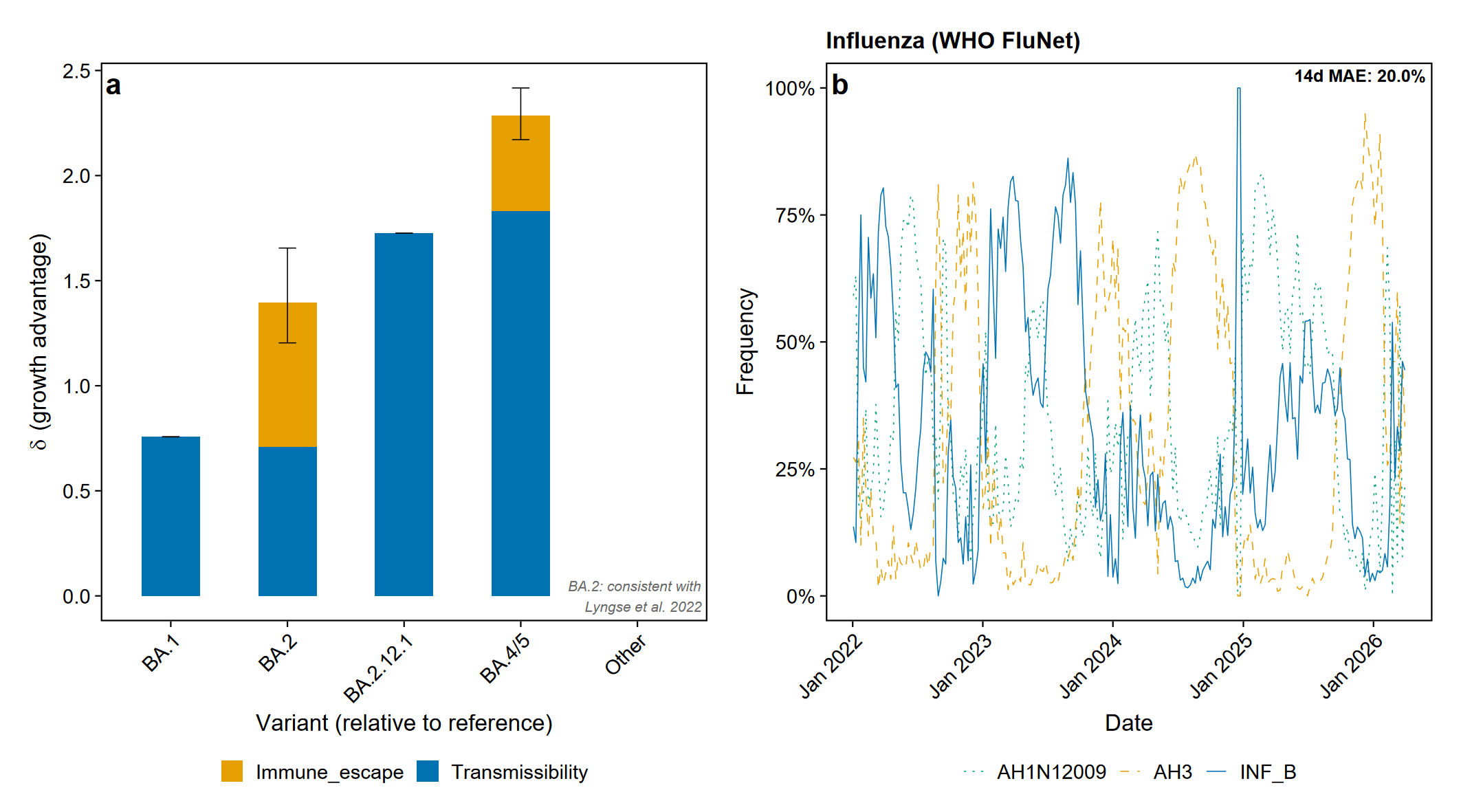
Relative Rt estimates are consistent with published values: BA.2 = 1.34x vs BA.1 (Lyngse et al. 2022, published 1.3–1.5x); KP.3 = 1.36x vs JN.1. Generation times: 3.2 days for Omicron BA.* subvariants (Du et al. 2022); 5.0 days for JN/KP lineages.
Validated on real data
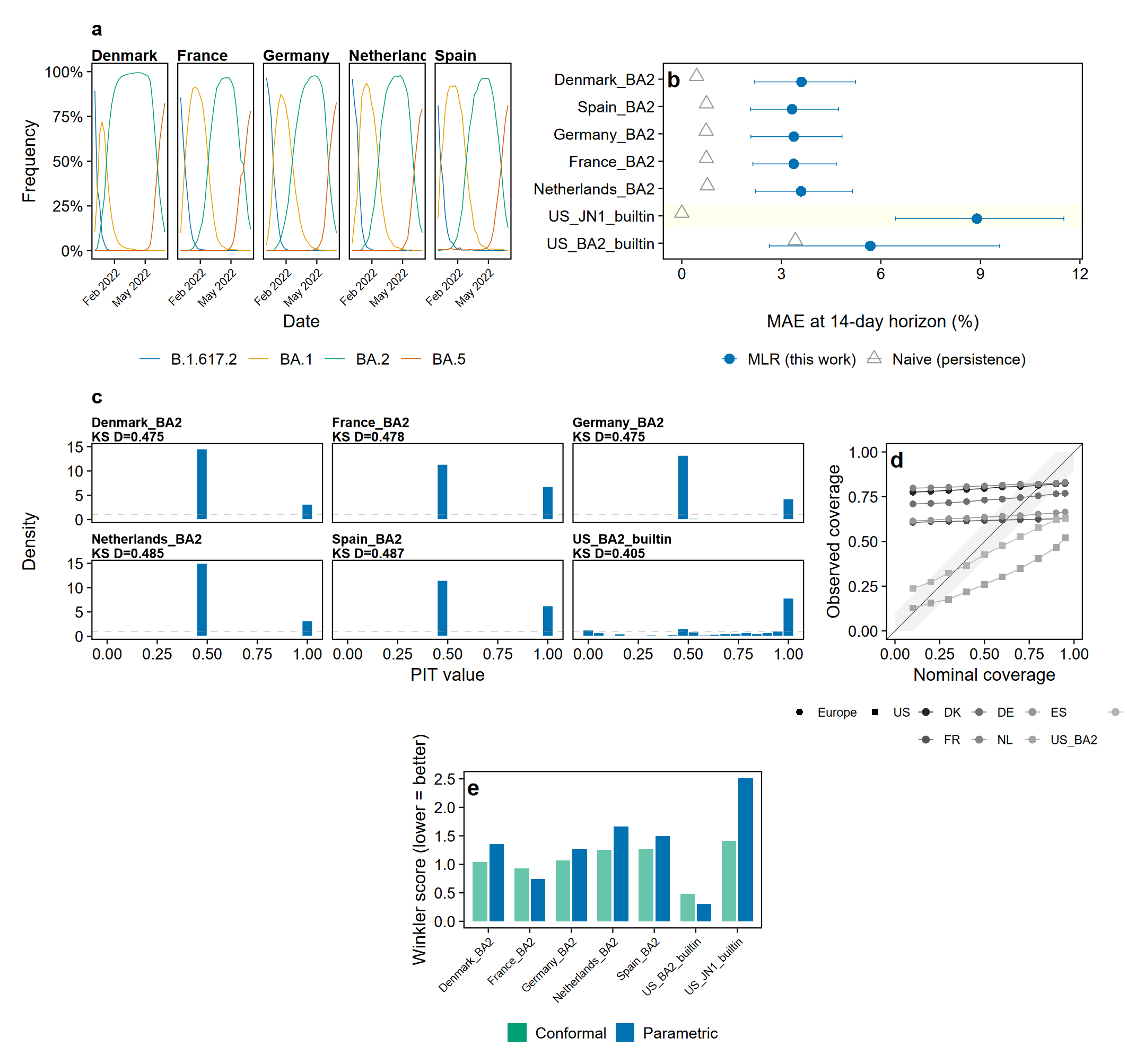
Rolling-origin evaluation across 7 datasets in 6 countries (5 European countries via ECDC + 2 US datasets via CDC):
| Dataset | 7d MAE | 14d MAE | 28d MAE |
|---|---|---|---|
| Denmark | 2.4 pp | 3.6 pp | 3.8 pp |
| France | 3.1 pp | 3.4 pp | 3.5 pp |
| Germany | 2.6 pp | 3.4 pp | 3.9 pp |
| Netherlands | 3.3 pp | 3.6 pp | 3.7 pp |
| Spain | 3.3 pp | 3.3 pp | 3.4 pp |
| US (BA.2) | 1.3 pp | 5.7 pp | 7.9 pp |
| US (JN.1) | — | 8.9 pp | 9.9 pp |
MAE in percentage points (pp). European data from ECDC (BA.2 period). US data from CDC (public domain). Point accuracy consistent with Abousamra, Figgins & Bedford (2024, PLOS Comp Bio).
Calibration diagnostics via calibrate() reveal that standard parametric prediction intervals are systematically underdispersed. Conformal prediction via conformal_forecast() provides correctly calibrated intervals. See analysis/ for reproducible validation scripts.
Features
Model fitting - fit_model() with engines "mlr", "hier_mlr", "piantham", "fga", "garw" (Bayesian engines require CmdStan)
Inference - Growth advantage in four scales: growth rate, relative Rt, selection coefficient, doubling time
Forecasting - Probabilistic frequency forecasts with parametric simulation and configurable sampling noise
Evaluation - Rolling-origin backtesting via backtest() with standardised scoring (MAE, RMSE, coverage, WIS) via score_forecasts()
Prediction calibration (v0.3.0+) - calibrate(): PIT histograms, reliability diagrams, KS uniformity test - recalibrate(): isotonic regression and Platt scaling - conformal_forecast(): distribution-free prediction intervals via split conformal inference and adaptive conformal inference - Proper scoring rules: CRPS, log score, DSS, calibration error
Immune-aware fitness estimation (v0.4.0+) - immune_landscape(): encode population immunity from seroprevalence, vaccination, or model-based data - fitness_decomposition(): partition growth advantage into intrinsic transmissibility vs immune escape - fit_dms_prior(): penalised MLR with Deep Mutational Scanning escape scores for early-emergence detection - selective_pressure(): genomics-only early warning signal
Surveillance optimisation (v0.5.0+) - surveillance_value(): Expected Value of Information for sequencing - adaptive_design(): Thompson sampling / UCB allocation across regions - detection_horizon(): weeks-to-detection under logistic growth - alert_threshold(): SPRT and CUSUM sequential detection - surveillance_dashboard(): multi-panel surveillance quality report
Surveillance utilities - summarize_emerging(): binomial GLM trend tests per lineage - sequencing_power(): minimum sample size for detection - collapse_lineages(), filter_sparse(): preprocessing
Visualisation - autoplot() methods for fits, forecasts, and backtest summaries - Colorblind-safe palettes
Interoperability - broom-compatible: tidy(), glance(), augment() - as_lfq_data() generic for extensible data import - read_lineage_counts() for CSV input
Supported pathogens
Any pathogen with variant/lineage-resolved sequencing count data: SARS-CoV-2, influenza, RSV, mpox, and others.
Citation
If you use lineagefreq in published work, please cite:
Gao C (2026). lineagefreq: Modelling Pathogen Lineage Frequency Dynamics and Forecasting Variant Replacement from Genomic Surveillance Data. R package version 0.5.1. https://CRAN.R-project.org/package=lineagefreq
citation("lineagefreq")