Overview
lineagefreq provides multiple modeling engines through the unified
fit_model() interface. This vignette shows how to compare
them using the built-in backtesting framework.
Three engines are available in v0.1.0:
- mlr: Multinomial logistic regression (frequentist, fast).
- piantham: Converts MLR growth rates to relative reproduction numbers using a generation-time approximation.
- hier_mlr: Hierarchical MLR with partial pooling across locations via empirical Bayes shrinkage.
Engine 1: MLR
The default engine fits a multinomial logistic regression with one growth rate parameter per non-reference lineage.
data(sarscov2_us_2022)
x <- lfq_data(sarscov2_us_2022,
lineage = variant, date = date,
count = count, total = total)
fit_mlr <- fit_model(x, engine = "mlr")
growth_advantage(fit_mlr, type = "growth_rate")
#> # A tibble: 5 × 6
#> lineage estimate lower upper type pivot
#> <chr> <dbl> <dbl> <dbl> <chr> <chr>
#> 1 BA.1 0 0 0 growth_rate BA.1
#> 2 BA.2 0.231 0.229 0.232 growth_rate BA.1
#> 3 BA.4/5 0.400 0.398 0.402 growth_rate BA.1
#> 4 BQ.1 0.352 0.346 0.357 growth_rate BA.1
#> 5 Other 0.151 0.149 0.152 growth_rate BA.1Engine 2: Piantham
The Piantham engine wraps MLR and translates growth rates to relative effective reproduction numbers using a specified mean generation time.
fit_pian <- fit_model(x, engine = "piantham",
generation_time = 5)
growth_advantage(fit_pian, type = "relative_Rt",
generation_time = 5)
#> # A tibble: 5 × 6
#> lineage estimate lower upper type pivot
#> <chr> <dbl> <dbl> <dbl> <chr> <chr>
#> 1 BA.1 1 1 1 relative_Rt BA.1
#> 2 BA.2 1.18 1.18 1.18 relative_Rt BA.1
#> 3 BA.4/5 1.33 1.33 1.33 relative_Rt BA.1
#> 4 BQ.1 1.29 1.28 1.29 relative_Rt BA.1
#> 5 Other 1.11 1.11 1.11 relative_Rt BA.1Comparing fit statistics
glance() returns a one-row summary for each model. Since
Piantham is a wrapper around MLR, the log-likelihood and AIC are
identical.
dplyr::bind_rows(
glance.lfq_fit(fit_mlr),
glance.lfq_fit(fit_pian)
)
#> # A tibble: 2 × 10
#> engine n_lineages n_timepoints nobs df logLik AIC BIC pivot
#> <chr> <int> <int> <int> <int> <dbl> <dbl> <dbl> <chr>
#> 1 mlr 5 40 461424 8 -465911. 931838. 931852. BA.1
#> 2 piantham 5 40 461424 8 -465911. 931838. 931852. BA.1
#> # ℹ 1 more variable: convergence <int>Backtesting
The backtest() function implements rolling-origin
evaluation. At each origin date, the model is fit on past data and
forecasts are compared to held-out future observations.
bt <- backtest(x,
engines = c("mlr", "piantham"),
horizons = c(7, 14, 21),
min_train = 56,
generation_time = 5
)
#> Backtesting ■■■ 7% | ETA: 14s
#> Backtesting ■■■■■■■■■ 26% | ETA: 11s
#> Backtesting ■■■■■■■■■■■■■■ 42% | ETA: 10s
#> Backtesting ■■■■■■■■■■■■■■■■■■ 55% | ETA: 8s
#> Backtesting ■■■■■■■■■■■■■■■■■■■■■■ 70% | ETA: 6s
#> Backtesting ■■■■■■■■■■■■■■■■■■■■■■■■■ 81% | ETA: 4s
#> Backtesting ■■■■■■■■■■■■■■■■■■■■■■■■■■■■■ 92% | ETA: 2s
#> Backtesting ■■■■■■■■■■■■■■■■■■■■■■■■■■■■■■■ 100% | ETA: 0s
bt
#>
#> ── Backtest results
#> 900 predictions across 31 origins
#> Engines: "mlr, piantham"
#> Horizons: 7, 14, 21 days
#>
#> # A tibble: 900 × 9
#> origin_date target_date horizon engine lineage predicted lower upper
#> * <date> <date> <int> <chr> <chr> <dbl> <dbl> <dbl>
#> 1 2022-03-05 2022-03-12 7 mlr BA.1 0.44 0.34 0.53
#> 2 2022-03-05 2022-03-12 7 mlr BA.2 0.18 0.11 0.26
#> 3 2022-03-05 2022-03-12 7 mlr BA.4/5 0 0 0.02
#> 4 2022-03-05 2022-03-12 7 mlr BQ.1 0 0 0.01
#> 5 2022-03-05 2022-03-12 7 mlr Other 0.37 0.28 0.47
#> 6 2022-03-05 2022-03-19 14 mlr BA.1 0.4 0.31 0.5
#> 7 2022-03-05 2022-03-19 14 mlr BA.2 0.21 0.13 0.29
#> 8 2022-03-05 2022-03-19 14 mlr BA.4/5 0 0 0.0252
#> 9 2022-03-05 2022-03-19 14 mlr BQ.1 0 0 0.01
#> 10 2022-03-05 2022-03-19 14 mlr Other 0.39 0.29 0.49
#> # ℹ 890 more rows
#> # ℹ 1 more variable: observed <dbl>Scoring
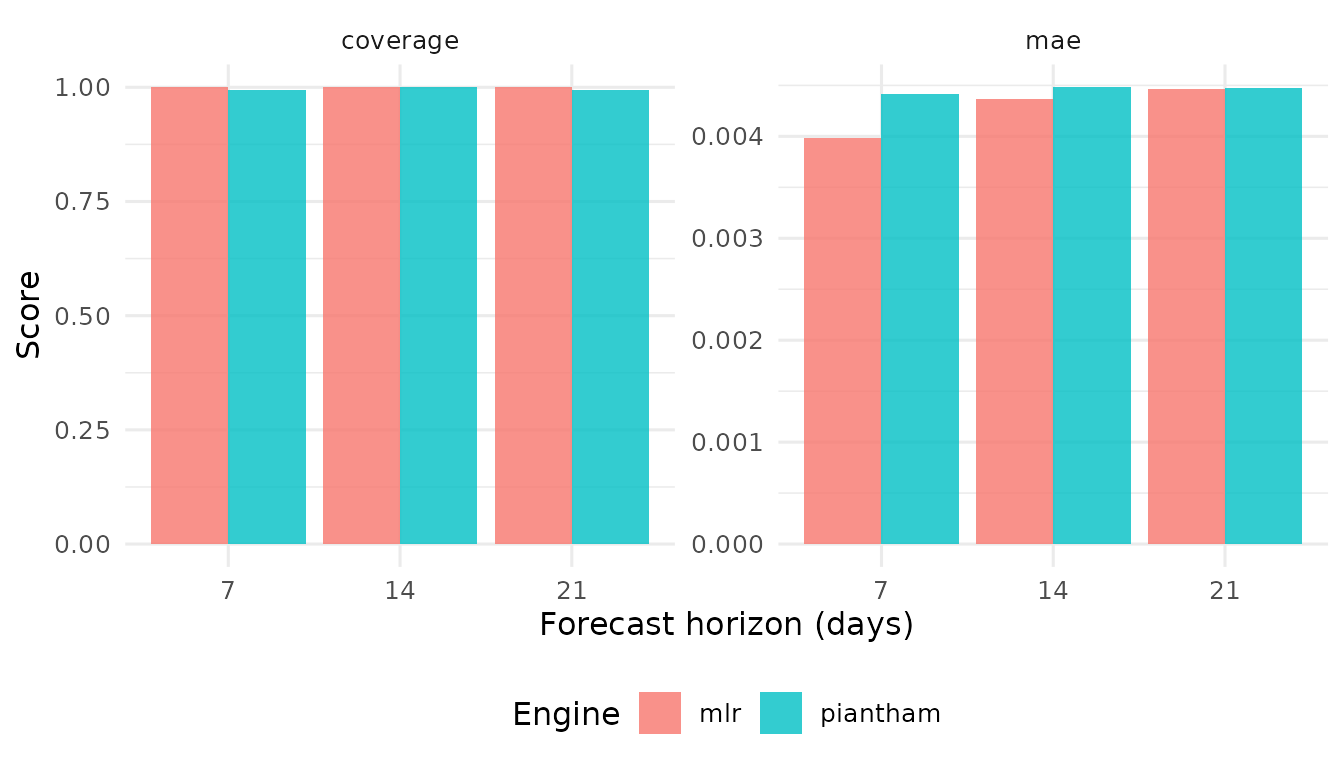
score_forecasts() computes standardized accuracy
metrics.
sc <- score_forecasts(bt,
metrics = c("mae", "coverage"))
sc
#> # A tibble: 12 × 4
#> engine horizon metric value
#> <chr> <int> <chr> <dbl>
#> 1 mlr 7 mae 0.00399
#> 2 mlr 7 coverage 1
#> 3 mlr 14 mae 0.00436
#> 4 mlr 14 coverage 1
#> 5 mlr 21 mae 0.00447
#> 6 mlr 21 coverage 1
#> 7 piantham 7 mae 0.00441
#> 8 piantham 7 coverage 0.994
#> 9 piantham 14 mae 0.00448
#> 10 piantham 14 coverage 1
#> 11 piantham 21 mae 0.00447
#> 12 piantham 21 coverage 0.993Model ranking
compare_models() summarizes scores per engine, sorted by
MAE.
compare_models(sc, by = c("engine", "horizon"))
#> # A tibble: 6 × 4
#> engine horizon mae coverage
#> <chr> <int> <dbl> <dbl>
#> 1 mlr 7 0.00399 1
#> 2 mlr 14 0.00436 1
#> 3 piantham 7 0.00441 0.994
#> 4 mlr 21 0.00447 1
#> 5 piantham 21 0.00447 0.993
#> 6 piantham 14 0.00448 1When to use which engine
| Scenario | Recommended engine |
|---|---|
| Single location, quick estimate | mlr |
| Need relative Rt interpretation | piantham |
| Multiple locations, sparse data | hier_mlr |
| Time-varying fitness (v0.2) | garw |
Hierarchical MLR
When data spans multiple locations with unequal sequencing depth,
hier_mlr shrinks location-specific estimates toward the
global mean. This stabilizes estimates for low-data locations.
A demonstration requires multi-location data, which the built-in
single-location dataset does not provide. See ?fit_model
for an example with simulated multi-location data.